Spotlight on Research: Top-Notch Researchers Propel Pitt's Drug Discovery Institute
Sitting atop the University of Pittsburgh’s Biomedical Science Tower 3 (BST3), researchers in Pitt’s Drug Discovery Institute (DDI) are searching for new methods to alleviate suffering caused by a variety of diseases. Their research includes studying the delicate protein interactions involved with cancer, designing new drug compounds, and developing automated technologies to enhance existing research methods. Their interests are diverse, yet their mission is the same—to advance global heath.
 Pitt’s Drug Discovery Institute has entered an exciting transition period under the new leadership of D. Lansing Taylor (right), Allegheny Foundation Professor of computational and systems biology in Pitt’s School of Medicine. The institute has four associate directors, including Barry Gold (left), a professor and chair of pharmaceutical sciences; Edward Chu, professor of medicine and of pharmacology and chemical biology as well as chief of the Pitt School of Medicine’s Division of Hematology-Oncology and deputy director of the University of Pittsburgh Cancer Institute; Peter Wipf, Distinguished University Professor of Chemistry and director of Pitt’s Combinatorial Chemistry Center; and Ivet Bahar, John K. Vries Professor and chair of computational and systems biology.
Pitt’s Drug Discovery Institute has entered an exciting transition period under the new leadership of D. Lansing Taylor (right), Allegheny Foundation Professor of computational and systems biology in Pitt’s School of Medicine. The institute has four associate directors, including Barry Gold (left), a professor and chair of pharmaceutical sciences; Edward Chu, professor of medicine and of pharmacology and chemical biology as well as chief of the Pitt School of Medicine’s Division of Hematology-Oncology and deputy director of the University of Pittsburgh Cancer Institute; Peter Wipf, Distinguished University Professor of Chemistry and director of Pitt’s Combinatorial Chemistry Center; and Ivet Bahar, John K. Vries Professor and chair of computational and systems biology.Drug discovery is a classic example of translational science, or “bench to bedside” research (or in this case, “bench to bottle”). Advances made in the laboratory lead to new and more effective pharmaceutical products for patients. The road to drug discovery, however, is never a straight path. In the early stages, medicinal chemists and structural biologists dissect the intricate interactions between proteins so they can hypothesize potential drug targets and determine which compounds are likely to disrupt or enhance a biological function. Once they’ve identified an area of interest, that target is screened against thousands of compounds to find one (or several) that might disrupt the protein-protein interaction. When a “hit” is identified, organic chemists find the best and most efficient methods to synthesize and scale up those compounds in the laboratory. Biologists then validate the compounds in the lab and in living things to determine effectiveness, safety, and stability. If any of these qualities is less than optimal, the molecule gets sent back to the chemists for improvement.
A Novel Concept
Pitt’s Drug Discovery Institute opened its doors in 2006, one of the first tenants of the $205 million BST3, a research tower with modern, open laboratories and a sophisticated floor plan. When Arthur S. Levine, senior vice chancellor for the health sciences and dean of Pitt’s School of Medicine, designed the concept for BST3, he envisioned collaboration at the most basic levels of science, believing that sharing physical space and technologies would encourage more collaboration. Barry Gold, DDI associate director and professor and chair of pharmaceutical sciences in the School of Pharmacy, thinks Levine’s concept works: “We really focus on teamwork here at Pitt. In fact, other universities are copying how we’re keeping our experts and their equipment in close proximity. I just wish the cancer center wasn’t so far away,” he laughs. (The University of Pittsburgh Cancer Institute is a five-minute bus ride from Pitt’s main campus.)
Over the years, DDI has expanded to a high-production facility, capable of holding as many as five million chemical compounds and equipped with more than 10 robots for automated assay plating, giving researchers virtually infinite drug- screening opportunities. Its faculty members hail primarily from three Pitt schools—the School of Arts and Sciences, the School of Medicine, and the School of Pharmacy—and create a unique mosaic of scientists, from organic chemists to clinical scientists, who work along the continuum of drug discovery.
Dennis Curran, a Pitt Distinguished Service and Bayer Professor of Chemistry in the School of Arts and Sciences, notes that chemists and clinicians often don’t speak the same language, but he believes the key to continuity is having leaders who can span the scientific spectrum. “Everyone has [his/her] own niche in academia, but we have great leaders who can relate to people up and down the pipeline,” says Curran, who specializes in organic compound synthesis.
The Research Team
Curran was a key collaborator in the development of AR67, an anticancer drug that is currently in phase II clinical trials with Arno Therapeutics Inc. AR67 is being tested in patients with glioblastoma multiforme, a highly aggressive brain cancer. He developed the drug with the late Thomas G. Burke, a professor of medicine at the University of Kentucky College of Medicine. Currently, Curran is working on the synthesis of several compounds: “Being academicians, we aren’t satisfied with just making more of the same compound. If we have a difficult synthesis, we want to make it better. We’re interested in pushing the edge of the possible in moving a compound to a drug candidate. We want new chemistry, new compounds, cutting-edge science.”
 Dennis Curran, a Pitt Distinguished Service Professor and Bayer Professor of Chemistry in the School of Arts and Sciences, was a key collaborator in the development of AR67, an anticancer drug that is currently in phase II clinical trials with Arno Therapeutics Inc. AR67 is being tested in patients with glioblastoma multiforme, a highly aggressive brain cancer.
Dennis Curran, a Pitt Distinguished Service Professor and Bayer Professor of Chemistry in the School of Arts and Sciences, was a key collaborator in the development of AR67, an anticancer drug that is currently in phase II clinical trials with Arno Therapeutics Inc. AR67 is being tested in patients with glioblastoma multiforme, a highly aggressive brain cancer.Alexander Dömling, professor of pharmaceutical sciences in the School of Pharmacy and of chemistry, came to Pitt in 2006 after founding two pharmaceutical companies and helping advance Almorexant, a sleep-aid drug, to phase III clinical trials. Despite his extensive industry experience, Dömling sees himself first as a teacher and researcher. “High throughput screening (HTS) is very important in drug discovery, but in academia, our priority is to teach. Before they use HTS, students need to understand the science behind drug interactions,” he says. Dömling’s main interest lies in designing antagonists of protein-protein interactions (PPI) that could be used as drag targets. He mines structural models to find small pockets of deeply buried amino acids, which he calls “anchors.” Together with collaborator Carlos Camacho, a professor of computational and systems biology in the School of Medicine, Dömling created an online tool, called anchor.query (http://anchorquery.ccbb.pitt.edu/), which allows researchers to search easily for these anchors and define promising areas for drug discovery. Dömling’s current PPI of interest is the interaction between the cancer-related proteins p53 and MDM2/MDMX. When the proteins interact, they form a small, deep cavity—the type that is ideal for drugs to bind to and effect a change in function. “This PPI is a hot topic in drug discovery right now, and no drugs have been approved that disrupt this type of interaction in cancer cells,” Dömling says. Dömling will be leaving Pitt in September to become chair of chemistry at the University of Groningen in The Netherlands, but will maintain research collaborations at Pitt via an adjunct appointment.
 Peter Wipf, Distinguished University Professor of Chemistry, was instrumental in founding Pitt’s Combinatorial Chemistry Center in 1998. The center was expanded in 2002 as part of a larger National Institutes of Health center initiative and became the University of Pittsburgh Center for Chemical Methodologies and Library Development. Wipf believes this type of diverse, multi-investigator center of excellence allows scholars to pool their expertise to discover novel therapies for major as well as neglected diseases.
Peter Wipf, Distinguished University Professor of Chemistry, was instrumental in founding Pitt’s Combinatorial Chemistry Center in 1998. The center was expanded in 2002 as part of a larger National Institutes of Health center initiative and became the University of Pittsburgh Center for Chemical Methodologies and Library Development. Wipf believes this type of diverse, multi-investigator center of excellence allows scholars to pool their expertise to discover novel therapies for major as well as neglected diseases. Peter Wipf is Distinguished University Professor of Chemistry, an associate director of DDI, and director of Pitt’s Combinatorial Chemistry Center, which was established in 1998. He is best known for his groundbreaking work in the total synthesis of complex organic molecules and was instrumental in advancing the use of combinatorial chemistry techniques at Pitt. Combinatorial chemistry is a synthesis method that became popular in the 1990s as a way for chemists to create large numbers of compounds in a very short period of time. As these new technologies and methods of organic synthesis were further developed at the Combinatorial Chemistry Center, the center was expanded in 2002 as part of a larger National Institutes of Health center initiative and became the University of Pittsburgh Center for Chemical Methodologies and Library Development (UPCMLD). UPCMLD is one of five centers in NIH’s Centers of Excellence in Chemical Methods and Library Development Program, which fosters innovation in chemical synthesis and drug discovery by developing chemical methodologies and creating diverse compound libraries with a range of physiological properties. As UPCMLD director, Wipf encourages interdisciplinary, multi-investigator research projects that allow the sharing of compounds, resources, and ideas. While Wipf is particularly interested in developing anticancer agents and neurodegenerative disease therapies, he feels that this type of diverse, multi-investigator center of excellence allows scholars to pool their expertise to discover novel therapies for major as well as neglected diseases.
Ivet Bahar is the John K. Vries Professor and chair of computational and systems biology and the newest associate director of DDI. Her research, like Dömling’s, focuses on protein-protein interactions, but she offers a unique biological perspective. She explains that her lab has traditionally looked at rational drug design as a dynamic process at the molecular level. By studying the atomic motions of proteins, researchers can glean how these proteins interact and rationalize which are likely to be inhibited by small molecules. “In the newest wave of drug discovery, however, we realize that you can design an inhibitor that binds well and has high activity, but we want to know what happens when you put it in the body,” Bahar says, adding that her group focuses on protein dynamics, or how the proteins move and interact. “Structure is important, but it is a snapshot. We want to observe how the protein achieves its function―then we can design the drug,” she adds.
 Ivet Bahar is the John K. Vries Professor and chair of computational and systems biology and an associate director of DDI. Her research team focuses on protein dynamics, or how proteins move and interact, in its efforts to design effective drugs.
Ivet Bahar is the John K. Vries Professor and chair of computational and systems biology and an associate director of DDI. Her research team focuses on protein dynamics, or how proteins move and interact, in its efforts to design effective drugs.Joining this systems biology approach is Andreas Vogt, a research assistant professor of computational and systems biology in the School of Medicine. Vogt has done substantial work in high-content imaging technology, which allows researchers to obtain data from individual cells and analyze the effects of drugs on their target of interest in a cellular context. Vogt also performs image-based analysis of whole organisms, such as zebra-fish, which are an ideal system for in vivo high-content screens. These tropical freshwater fish are vertebrates and share biological similarities to higher organisms; they are optically transparent, and their tiny embryos can easily be stored in 96-well microplates, a standard format for high throughput screening. Using his imaging technology and an artificial intelligence-based image analysis method called Cognition Network Therapy, Vogt has detected and quantified structures of interest in zebrafish embryos, including the presence of intersegmental blood vessels, which are used as a measure of angiogenesis. Vogt collaborates with Pitt’s zebrafish superstars, professors of developmental biology Michael Tsang and Neil Hukriede, in developing novel automated imaging assays to measure fluorescence and analyze biological function in transgenic zebrafish embryos.
Pitt researchers employ both HTS, the dominant tool in the drug discovery process, and high- content screening (HCS)—a combination of fluorescence labeling technologies and electronic imaging technologies that enable the study of individual cells on a light microscope. Together, those tools allow collaborators to access Pitt’s massive compound library, where researchers can screen thousands of interesting chemotypes [a chemically distinct entity in a plant or microorganism] against their target of interest and visualize protein interactions via live cell microscopy. Paul Johnston is a research professor of pharmaceutical sciences and manager of the Pittsburgh Molecular Library Screening Center (PMLSC), a federally funded joint venture between Pitt and Carnegie Mellon University that was part of the NIH Roadmap Initiative Molecular Library Screening Center Network pilot phase. Johnston is also coprincipal investigator and manager of the Pittsburgh Specialized Application Center (PSAC), a component of the National Cancer Institute’s Chemical Biology Consortium (CBC). CBC is part of the Experimental Therapeutics (NExT) Program, a partnership between the National Cancer Institute’s Division of Cancer Treatment and Diagnosis and Center for Cancer Research, and is tasked with streamlining the development and testing of promising new anticancer drugs to expedite their delivery to the bedside.
Johnston is a pioneer in developing and performing high throughput cell-based screening assays. He worked in the pharmaceutical industry for 14 years and has close ties with Pittsburgh biotechnology experts. This experience, along with his creativity and innovation with HTS and HCS assays, has allowed him to build one of the nation’s top academic screening facilities. The PMLSC and PSAC offer advanced services to investigators who are interested in screening large libraries of small molecules against targets of interest, using such advanced technologies as liquid-handling robots; multimode plate readers with luminescence, fluorescence, fluorescence polarization, time-resolved fluorescence, and absorbance detection capability; HCS fluorescent microscopy platforms with live cell and kinetic screening options with image storage and analysis databases; sophisticated screening microscopes; and biosafety level 2 tissue culture hoods and incubators.
Combining Business and Basic Science
DDI is in an exciting transition period, as it shifts focus with its new director D. Lansing Taylor, Allegheny Foundation professor in the School of Medicine’s Department of Computational and Systems Biology. Taylor pioneered high content screening methods to automate cell and experimental animal drug discovery and marketed these technologies through his company, Cellomics, Inc., now part of ThermoFisher. He also founded several Pittsburgh-based biotechnology companies, most recently Cellumen and Cernostics, which applied cellular and tissue systems biology to drug safety and diagnostics.
As both an entrepreneur and an academician, Taylor provides a unique industrial perspective with a deep appreciation for basic science. “There is no single isolated target that we can perturb with drugs, so it is essential to understand the biological systems and address this complexity up front, using functional model systems such as sophisticated cell models and small experimental animal models, like C. elegans [roundworm] and zebrafish. We can use these live models to gain biological insight very early on in the drug discovery process.” Taylor is energetic about his new role and the future of drug discovery. Even before he began his first day in his new position, Taylor coined this tagline that demonstrates his enthusiasm and vision for DDI: “Novel chemistries and systems biologies to accelerate drug discovery.’”
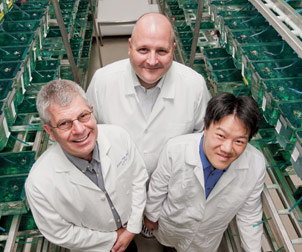 Andreas Vogt (left), a research assistant professor of computational and systems biology in Pitt’s School of Medicine, has done substantial work in high-content imaging technology, which allows researchers to obtain data from individual cells and analyze the effects of drugs on their target of interest in a cellular context. Vogt also performs image-based analysis of whole organisms, such as zebrafish. Because of their small size, optical transparency, and physiological similarities to human biology, zebrafish embryos are an ideal system for in vivo high-content screens. Vogt collaborates with Pitt Professors of Developmental Biology Neil Hukriede and Michael Tsang (center and right, respectively) in developing novel automated imaging assays to analyze biological function in transgenic fluorescent zebrafish embryos.
Andreas Vogt (left), a research assistant professor of computational and systems biology in Pitt’s School of Medicine, has done substantial work in high-content imaging technology, which allows researchers to obtain data from individual cells and analyze the effects of drugs on their target of interest in a cellular context. Vogt also performs image-based analysis of whole organisms, such as zebrafish. Because of their small size, optical transparency, and physiological similarities to human biology, zebrafish embryos are an ideal system for in vivo high-content screens. Vogt collaborates with Pitt Professors of Developmental Biology Neil Hukriede and Michael Tsang (center and right, respectively) in developing novel automated imaging assays to analyze biological function in transgenic fluorescent zebrafish embryos.Taylor plans to use his business acumen to reach out to potential partners in industry. He notes that both Harvard University and Washington University in St. Louis, two national leaders in drug discovery, have recently developed partnerships with large pharmaceutical companies. He is confident that Pitt’s academic prowess will attract the pharmaceutical industry, as well: “We have tremendous talent, world-class facilities, and the drive to push discovery into the future,” Taylor says. “By combining our intellectual power with their business knowledge, we will gain mutual value that will help our institution and the world. It’s all here.”
Associate Director Gold explains, “Drug discovery is a lengthy, difficult, expensive process. An entire drug portfolio from conception to delivery can cost upwards of $800 million, money that universities typically do not have. At the same time, the big pharma pipeline is not always full, so when that occurs (for example, when they have several drugs go off patent at the same time), they look to universities for real leads on new compounds.” Gold thinks the strength of the University is in studying a diverse set of targets and creating novel candidates for drug screening. He says, “The more work we do here, the less risk is involved for pharma, and they are more likely to purchase a license. Since we don’t have the capacity for large scale-ups or human trials, this is an ideal situation for the inventor, the University, and thereby the region as a whole.”
DDI looks to industry partnerships to help advance the basic research that might lead to new therapies and help society. “The money that comes with an industry partnership is reinvested into the University and supports additional research and infrastructure,” Gold says. This investment has clearly paid off, as Pitt has recruited some of the nation’s top scientists and has remained in the top 10 schools in the nation for NIH funding for more than 10 years.
Curran believes that this upward swing will continue into the next decade: “Our chemistry department is in the process of leapfrogging to the top with our new organic chemistry wing and $2 million NMR [nuclear magnetic resonance] facility that we use to characterize drug candidates. Faculty recruits are looking for three things in a university—a good name, high-quality faculty, and high-tech facilities―and we have all three. Good faculty attract good students, and funding typically follows.” Pitt’s remarkable success in the past 10 years should prove, without a doubt, that Curran’s belief is true.
Other Stories From This Issue
On the Freedom Road

Follow a group of Pitt students on the Returning to the Roots of Civil Rights bus tour, a nine-day, 2,300-mile journey crisscrossing five states.
Day 1: The Awakening
Day 2: Deep Impressions
Day 3: Music, Montgomery, and More
Day 4: Looking Back, Looking Forward
Day 5: Learning to Remember
Day 6: The Mountaintop
Day 7: Slavery and Beyond
Day 8: Lessons to Bring Home
Day 9: Final Lessons

